MethScope is an R package for ultra-fast analysis of sparse DNA methylome data using Most Recurrent Methylation Patterns (MRMPs).
It supports downstream analysis for: - Cell type annotation - Cell type deconvolution - Unsupervised clustering - Cancer cell-of-origin prediction - Missing value imputation
Why MethScope?
Sparse single-cell and spatial methylome data are often too sparse to analyze directly. MethScope compresses methylation signals into MRMP-based embeddings so you can run robust and scalable downstream tasks with standard analysis workflows.
Method overview
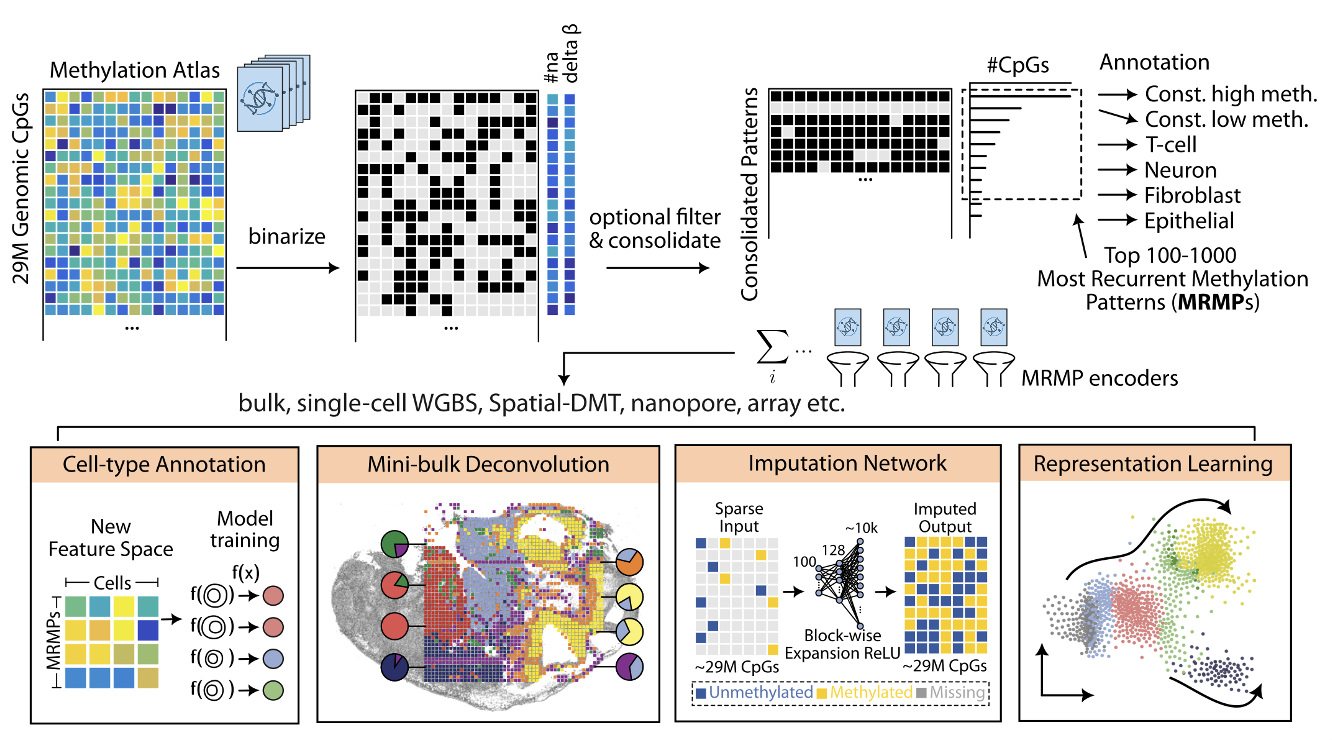
MethScope converts high-dimensional methylation atlas signals into compact MRMP features and applies these features across multiple analysis tasks.
Core workflow: - Binarize methylation atlas profiles and consolidate recurrent patterns - Select top recurrent methylation patterns (MRMPs) - Encode each sample, cell, or pixel into an MRMP-based representation - Run downstream modeling for annotation, deconvolution, imputation, and representation learning
Use cases supported in the current pipeline: - Cell-type annotation in sparse single-cell methylome profiles - Mini-bulk deconvolution for mixed-cell samples - Missing-value imputation for sparse CpG measurements - Representation learning for clustering and embedding analysis
Installation
Install from CRAN:
install.packages("MethScope")Or install the development version from GitHub:
remotes::install_github("zhou-lab/MethScope")Quick start
library(MethScope)
# 1) Generate MRMP embedding from your .cg file and MRMP reference
example_file <- "example.cg"
reference_pattern <- "Liu2021_MouseBrain.cm"
input_pattern <- GenerateInput(example_file, reference_pattern)
# 2) Predict cell types with a built-in model
pred <- PredictCellType(MethScope:::Liu2021_MouseBrain_P1000, input_pattern)
# 3) Visualize prediction results
PlotUMAP(input_pattern, pred)Tutorials and documentation
- Documentation website: zhou-lab.github.io/MethScope
- End-to-end tutorial: MethScope-Tutorial
- Building MRMP references: MethScope-MRMP
Agent skill
This repository includes a reusable MethScope agent skill under agent-skills/methscope/.
Codex
Install the skill under $CODEX_HOME/skills/methscope/ with this layout:
$CODEX_HOME/skills/methscope/
SKILL.md <- copy from agent-skills/methscope/codex/SKILL.md
core/Copy: - agent-skills/methscope/codex/SKILL.md to $CODEX_HOME/skills/methscope/SKILL.md - agent-skills/methscope/core/ to $CODEX_HOME/skills/methscope/core/
Then invoke the skill when working on MethScope package usage, vignettes, .cg and .cm inputs, MRMP embeddings, prediction, training, deconvolution, or visualization.
Claude
If you already keep a repository-level CLAUDE.md, copy the contents of agent-skills/methscope/claude/CLAUDE.md into it or reference that file from your existing Claude project instructions.
If you do not already have a project CLAUDE.md, use agent-skills/methscope/claude/CLAUDE.md as the starting project context for this repository.
Keep agent-skills/methscope/core/ in the repository, because the Claude instructions point to those shared files.
Data resources
- Example and reference data: zhou-lab/methscope_data
-
.cggeneration and preprocessing: YAME - Pattern interpretation: knowYourCG